Motivation
Foundation-scale sequence-to-function models have rapidly advanced regulatory genomics. Architectures like AlphaGenome and Enformer predict thousands of regulatory tracks across large genomic contexts and achieve impressive genome-wide accuracy (hence the term generalists).
Side note: sequence-to-function (seq2func) models learn a direct mapping from DNA sequence to one or more experimentally measured molecular readouts from assays such as chromatin accessibility, transcription factor binding, or gene expression.
These models also just continue to increase in their number of parameters, receptive fields and number of tasks they predict - if you’re skeptical just look at a selection of these recent models:
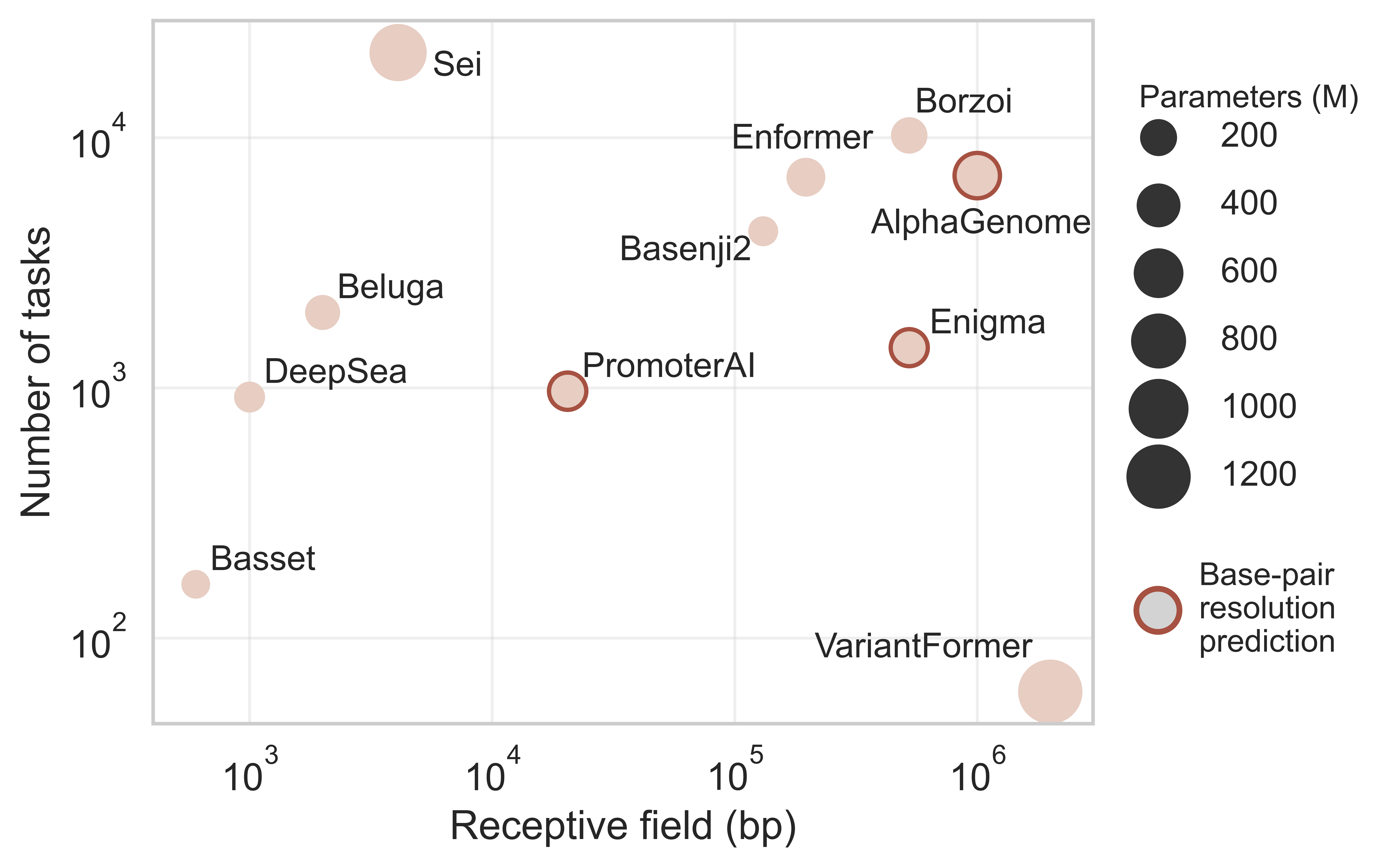
But many real experimental workflows don’t look like the genome. Perturbation assays — including MPRAs, enhancer design screens, and synthetic element optimisation — evaluate short (~100–300 bp) sequences outside their native context. Applying these now megabase-scale predictors to such data introduces unnecessary padding, compute overhead, and arbitrary flanking sequence assumptions which are just unsatisfactory!
We asked a simple question:
What if we treated these models as reusable regulatory feature extractors instead of end-to-end predictors?
The key idea: modular regulatory encoders
Modern seq2func models like AlphaGenome can be decomposed into three functional components:
Sequence encoder - learns motifs, spacing rules, and local regulatory syntax (e.g. convolutions and pooling)
Long-range context module - (e.g. transformers) models distal regulatory dependencies
Task decoder - predicts assay-specific outputs
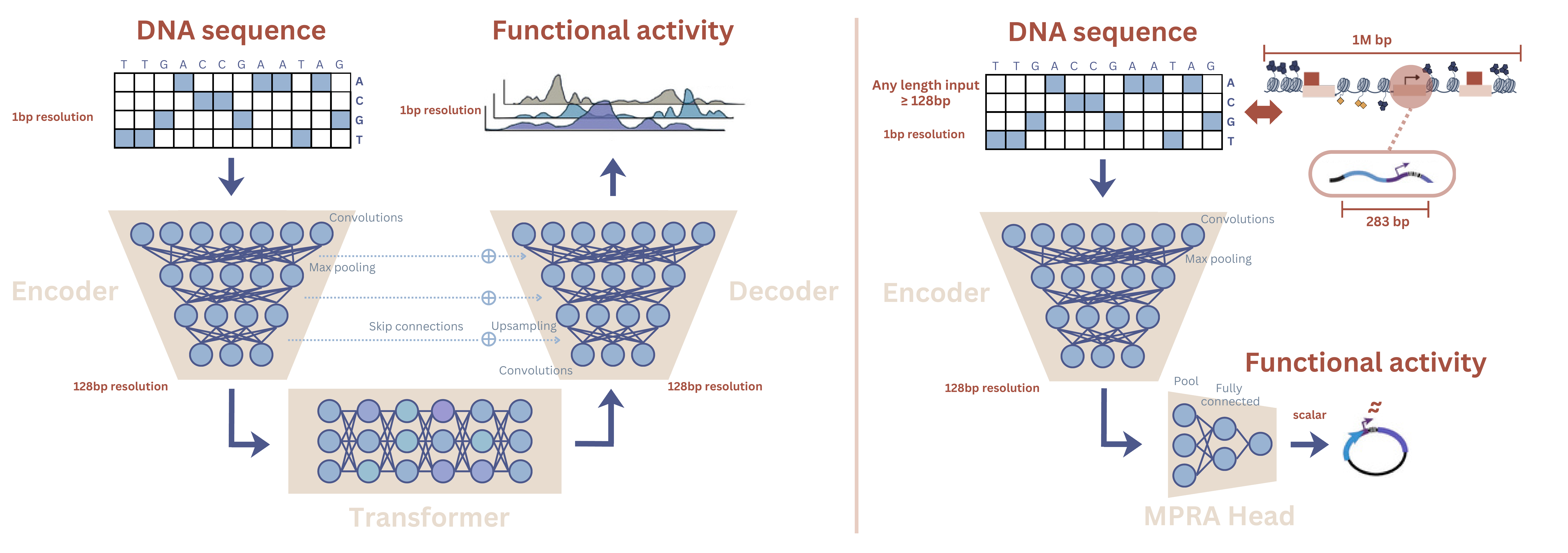
For short perturbation sequences assayed in isolation — such as MPRA constructs that test cis-regulatory activity outside their native chromosomal context — long-range genomic interactions are largely absent, so distal context modeling is often unnecessary. The encoder, however, contains rich regulatory representations learned from genome-scale supervision. We extract and reuse this encoder - see the image below:
Encoder intuition - In these models, the encoder progressively downsamples the input sequence through convolution and pooling operations, similar to how image CNNs compress spatial resolution while increasing feature richness. As a result, the encoder outputs a sequence of embeddings where each position summarises regulatory features over a window of roughly ~128 bp rather than single nucleotides. This resolution is sufficient to capture motif combinations and local regulatory syntax while keeping representations compact and computationally efficient.
Although AlphaGenome was trained on ~1 megabase genomic windows, we show that its convolutional encoder can be repurposed for much shorter sequences. This reflects a division of labor within the architecture: the encoder captures local regulatory grammar, while the transformer and decoder handle long-range integration and base-resolution track prediction. By isolating the encoder, we retain the reusable representation module while discarding machinery designed for distal genomic context — precisely the setting of MPRA assays and other tasks centered on local regulatory activity, such as chromatin accessibility prediction.
What we do:
isolate the convolutional encoder
adapt positional handling for short inputs
pool encoder embeddings
attach a lightweight regression head
optionally fine-tune or keep encoder frozen
This allows direct training on short sequences while preserving pretrained regulatory features! We applied this to AlphaGenome and Enformer (the later to highlight the generalisation of the approach).
Why this helps
Practical advantages:
supports variable-length inputs
removes megabase padding overhead
standardises comparisons across architectures
dramatically reduces inference cost — in our testing it was 500 fold quicker to run the encoder model than full AlphaGenome
Conceptual advantage:
- separates regulatory representation learning from task-specific prediction
Performance on MPRA and STARR-seq
Before I get into the how of doing this, let me convince you that it’s worthwhile — we evaluated modular encoders on:
Results:
achieved state-of-the-art accuracy on both tasks (subplots a-b below)
AlphaGenome encoder probing remained strong across species
Enformer benefited more from fine-tuning — perhaps its encoder learned less cis-regulatory logic
AlphaGenome required minimal adaptation as pretrained encoder already captures transferable signal
Side note: probing means the AlphaGenome encoder is frozen and only the added head is updated whereas fine-tuning means everything is updated (encoder and head).
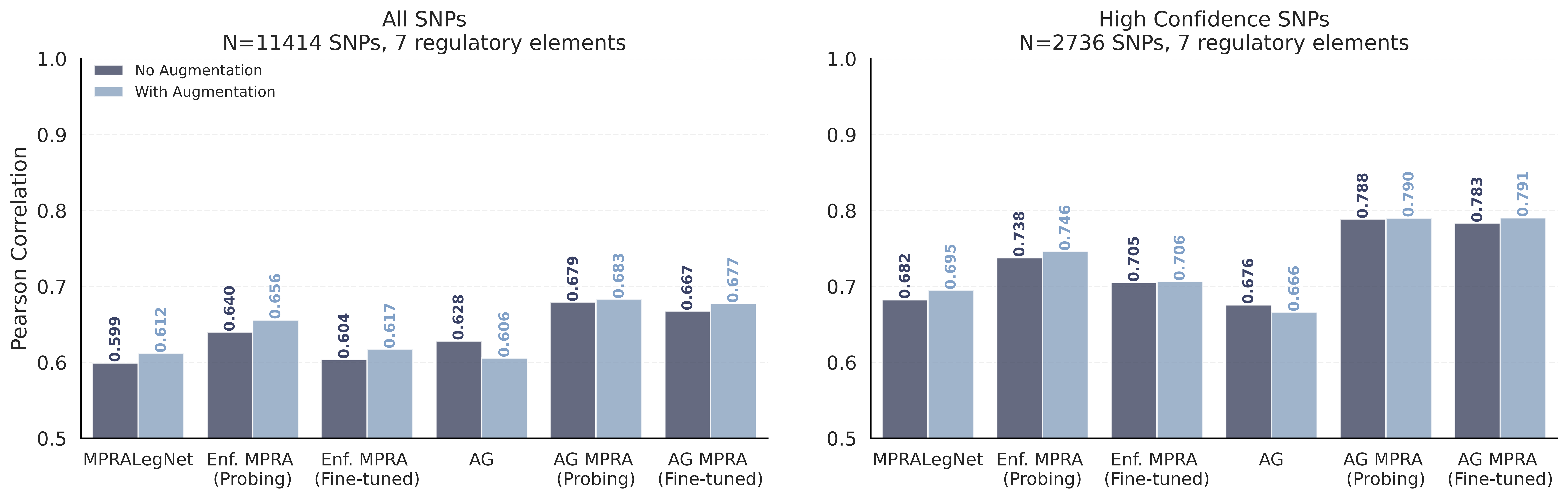
This supports the idea that genome-scale training learns reusable regulatory structure. The performance results:
, [DeepSTARR](https://www.nature.com/articles/s41588-022-01048-5), [DREAM-RNN](https://www.nature.com/articles/s41587-024-02414-w), and AlphaGenome (AG). We applied encoder extraction and fine-tuning to Enformer (Enf. MPRA) and AlphaGenome (AG MPRA), evaluated with probing (head-only) or encoder fine-tuning. Benchmark on lentiMPRA and STARR-seq](/blogs/2026-002/lenti_starr_res.png)
What matters when adapting encoders?
So in an attempt to push performance as much as possible, we did a hyperparameter sweep which revealed the:
Most important choices
deeper MLP heads
flattening encoder embeddings
Less important choices
optimiser choice
weight decay
learning rate schedule
Progressive unfreezing provided modest gains, with a slight benefit from delaying encoder updates. The results of this sweep is at the end of the post.
Transfer to regulatory variant prediction (CAGI5)
We next evaluated all models on the CAGI5 benchmark which provides experimentally measured effects of thousands of regulatory variants, making it a standard test for evaluating how well models predict functional impacts beyond the training assay.
Key findings
MPRA fine-tuning improved performance (using matched cell types with lentiMPRA models)
frozen encoder probing generalised better out-of-distribution
task-specific fine-tuning can introduce assay bias — full fine-tuning rather than probing led to the models overfitting on the lentiMPRA data and thus worse performance on the CAGI5 data.
A smaller aggregation window improved pretrained AlphaGenome’s performance (more on this below).
This may highlight a trade-off of specialisation vs generalisation, or with better regularisation maybe this could be controlled even with the larger number of free parameters. The results:
A Technical Note: Improving Base AlphaGenome’s Performance on CAGI5
When we tested against the AlphaGenome model before any fine-tuning on MPRA data, we noticed something interesting.
Aligning the aggregated window size to match the size of the MPRA assay (central 384 base-pairs) improved zero-shot prediction relative to AlphaGenome’s original protocol (central 501bp) by 25%!
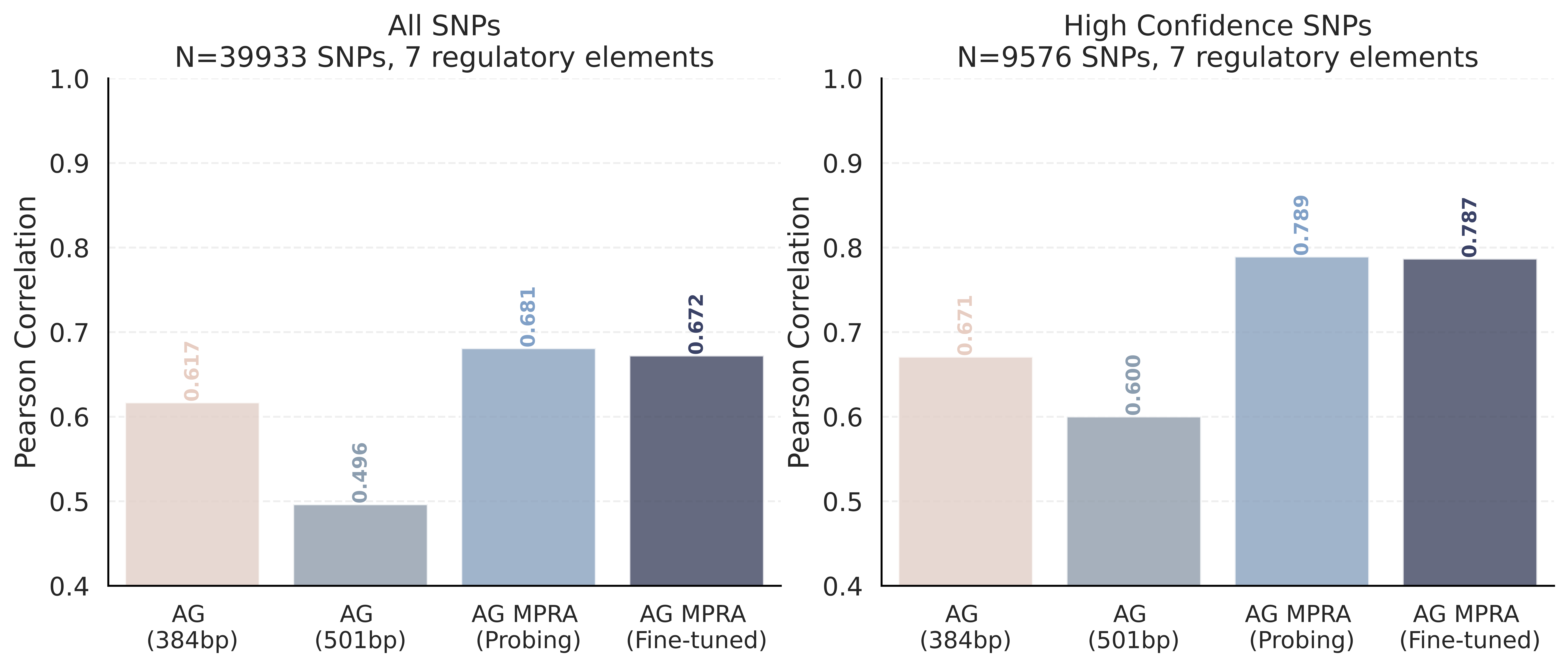
So I would advise testing differing aggregation windows if you are using AlphaGenome in this manner. Or, just use our extracted encoder approach which boosted performance by another 10%!
What transfers — and why?
So we should probably now take a step back, what are our results showing?
They highlight that encoder representations learned under genome-scale multitask supervision retain regulatory signal that transfers across:
assays
perturbation regimes
species (STARR-seq data was in fly, AlphaGenome was trained on human and mouse — this is pretty cool!)
This transfer was observed across distinct architectures (AlphaGenome and Enformer), suggesting that the modular encoder perspective is broadly applicable.
Implications for regulatory design workflows
Now to the so what? Well, encoder-only predictors have numerous advantages over their generalist parents, they enable:
rapid scoring of candidate constructs
iterative design → score → optimise loops
compute-efficient large-scale screening
Seq2func foundation models can therefore function as reusable regulatory representation engines inside perturbation pipelines — think of synthetic biology DNA design, where these models could help accelerate synthetic enhancer and promoter development (see this work for example).
Open questions
So what didn’t we explore here:
Which encoder layers contribute most to transfer?
How stable are representations across assays and species?
Can modular encoders accelerate generative regulatory design?
All of these would be really interesting future directions.
Takeaway — the TL;DR
Foundation seq2func models are typically used as monolithic predictors.
A modular view reveals something more useful:
Their encoders are transferable regulatory representation modules.
Extracting and adapting these representations enables efficient perturbation modeling, fair cross-model comparison, and scalable regulatory design workflows.
Code
Finally, how can you use this approach:
This analysis uses the native jax/haiku AlphaGenome wrapper package which is available from the Genomics x AI community github (see our post on this) and all code to run the analysis is here.
But here is a minimum script or if you would prefer to run it yourself on lentiMPRA data, see our colab notebook:
Tutorial
1. Model initialisation
from alphagenome.models import dna_output
from alphagenome_ft import (
templates,
CustomHeadConfig,
CustomHeadType,
register_custom_head,
create_model_with_heads,
)
# 1. Register an encoder-only head
register_custom_head(
"mpra_head",
templates.EncoderOnlyHead,
CustomHeadConfig(
type=CustomHeadType.GENOME_TRACKS,
output_type=dna_output.OutputType.RNA_SEQ,
num_tracks=1,
),
)
# 2. Create a model that uses encoder output only
model = create_model_with_heads(
"all_folds",
heads=["mpra_head"],
use_encoder_output=True, # ← CRITICAL for encoder-only mode
)
# 3. Optionally freeze backbone to start with heads-only finetuning
model.freeze_except_head("mpra_head")Key points:
use_encoder_output=Truebypasses the transformer/decoder stack and exposes encoder features at ~128 bp resolutiontemplates.EncoderOnlyHeadapplies a simple MLP on top of these embeddings
2. Training Loop
For MPRA-like data, you will typically have short sequences and scalar or low-dimensional outputs (e.g. log expression).
You can either:
- Use your own data loader and a custom training loop with
model.create_loss_fn_for_head, or - Follow the more complete MPRA scripts in the external repository.
Minimal example with a custom loop:
import jax
import jax.numpy as jnp
import optax
from alphagenome_ft import CustomHead
# Suppose you have: sequences_onehot: (B, L, 4), targets: (B, 1)
loss_fn = model.create_loss_fn_for_head("mpra_head")
optimizer = optax.adamw(learning_rate=1e-3, weight_decay=1e-4)
opt_state = optimizer.init(model._params)
def train_step(params, state, opt_state, batch_sequences, batch_targets):
def loss_inner(current_params):
preds_dict = model._predict(
current_params,
state,
batch_sequences,
jnp.zeros((batch_sequences.shape[0],), dtype=jnp.int32), # organism_index
negative_strand_mask=jnp.zeros((batch_sequences.shape[0],),
dtype=bool),
strand_reindexing=model._metadata[
next(iter(model._metadata))].strand_reindexing,
)
preds = preds_dict["mpra_head"]
loss_dict = loss_fn(
preds,
{"targets": batch_targets, "organism_index": None},
)
return loss_dict["loss"]
loss, grads = jax.value_and_grad(loss_inner)(params)
updates, new_opt_state = optimizer.update(grads, opt_state)
new_params = optax.apply_updates(params, updates)
return new_params, new_opt_state, lossConclusion — bridging genome-scale models and perturbation assays
Foundation sequence-to-function models are built for megabase context and genome-wide prediction. We show that their most transferable asset is much smaller: the convolutional encoder that learns cis-regulatory grammar.
By isolating this module, we:
repurpose genome-scale pretrained representations for 100–300 bp perturbation sequences
eliminate unnecessary long-range context machinery in assays that isolate regulatory elements
achieve state-of-the-art MPRA, STARR-seq and CAGI5 benchmark performance
reduce inference cost by orders of magnitude
generalise the approach across architectures
Despite being trained on ~1 Mb inputs, the AlphaGenome encoder adapts cleanly to >=128 bp sequences — matching the scale at which cis-regulatory logic operates in perturbation assays.
This reframes foundation genomics models not as monolithic predictors, but as modular regulatory representation engines that can be embedded directly into perturbation, design, and variant-effect workflows.
Code
Implementation and reproducible experiments: https://github.com/Al-Murphy/alphagenome_FT_MPRA
AlphaGenome encoder fine-tuning utilities: https://github.com/genomicsxai/alphagenome_ft
Hyperparameter sweep results
Stage 1
Stage 1 was a hyperparameter sweep for lentiMPRA with a frozen encoder (probing regime).
We varied the prediction head architecture and training hyperparameters while keeping encoder weights fixed. Note no reverse complement or random shift augementations were used for this benchmark. mlp-X-Y denotes a two-layer multilayer perceptron head with hidden dimensions X and Y; mlp-X denotes a single hidden layer of size X; pool-flatten uses global pooling followed by flattening; pool-center extracts the central token representation; do-p indicates dropout rate p applied to the head; wd-1eK indicates weight decay of $10^{-K}$; lr-plateau and lr-cosine denote ReduceLROnPlateau and cosine annealing learning rate schedules, respectively; opt-adamw indicates the AdamW optimiser; act-gelu replaces the default activation with GELU. Baseline used a single multilayer perceptron head of size 1024 with sum pooling, Adam optimiser and RELU activation, and no dropout, weight decay or learning rate plateau. Performance is reported as Pearson correlation on the held-out test fold for HepG2, K562, and WTC11, with average performance and rank across cell types.
| Hyperparameter | HepG2 | K562 | WTC11 | Average | Rank |
|---|---|---|---|---|---|
| mlp-512-512 | 0.8581 | 0.8258 | 0.7825 | 0.8221 | 1 |
| mlp-512-256 | 0.8573 | 0.8273 | 0.7803 | 0.8216 | 2 |
| pool-flatten | 0.8547 | 0.8250 | 0.7837 | 0.8211 | 3 |
| mlp-256-256 | 0.8544 | 0.8253 | 0.7814 | 0.8204 | 4 |
| mlp-128 | 0.8532 | 0.8235 | 0.7786 | 0.8185 | 5 |
| mlp-256 | 0.8527 | 0.8212 | 0.7758 | 0.8166 | 6 |
| pool-center | 0.8498 | 0.8211 | 0.7778 | 0.8162 | 7 |
| do-0.1 | 0.8521 | 0.8193 | 0.7755 | 0.8156 | 8 |
| do-0.2 | 0.8522 | 0.8188 | 0.7755 | 0.8155 | 9 |
| mlp-512 | 0.8514 | 0.8199 | 0.7752 | 0.8155 | 10 |
| do-0.5 | 0.8521 | 0.8188 | 0.7755 | 0.8155 | 11 |
| do-0.4 | 0.8521 | 0.8188 | 0.7755 | 0.8155 | 12 |
| do-0.3 | 0.8521 | 0.8188 | 0.7752 | 0.8154 | 13 |
| wd-1e6 | 0.8529 | 0.8171 | 0.7738 | 0.8146 | 14 |
| wd-1e5 | 0.8530 | 0.8166 | 0.7742 | 0.8146 | 15 |
| lr-plateau | 0.8526 | 0.8170 | 0.7733 | 0.8143 | 16 |
| ————– | ———- | ———- | —— | ———- | —- |
| baseline | 0.8526 | 0.8170 | 0.7733 | 0.8143 | 17 |
| ————– | ———- | ———- | —— | ———- | —- |
| opt-adamw | 0.8526 | 0.8161 | 0.7738 | 0.8142 | 18 |
| wd-1e4 | 0.8522 | 0.8167 | 0.7732 | 0.8141 | 19 |
| act-gelu | 0.8513 | 0.8167 | 0.7724 | 0.8134 | 20 |
| lr-cosine | 0.8399 | 0.8007 | 0.7605 | 0.8004 | 21 |
Stage 2
Stage 2 was a hyperparameter sweep for lentiMPRA with encoder unfreezing (fine-tuning regime).
Starting from the best Stage 1 configuration, we varied the unfreezing schedule. s2-s1epN denotes unfreezing the encoder after N epochs of head-only training; s2-baseline denotes the default unfreezing schedule used in the main experiments (unfreezing triggered by validation loss plateau). Baseline used a single multilayer perceptron head of size 1024 with sum pooling, Adam optimiser and RELU activation, and no dropout, weight decay or learning rate plateau. All models used reverse complement and random shift augmentations. Performance is reported as Pearson correlation on the held-out test fold for HepG2, K562, and WTC11, with average performance and rank across cell types.
| Hyperparameter | HepG2 | K562 | WTC11 | Average | Rank |
|---|---|---|---|---|---|
| s2-s1ep3 | 0.8663 | 0.8439 | 0.7730 | 0.8277 | 1 |
| ————— | ———- | ———- | ———- | ———- | —- |
| s2-baseline | 0.8701 | 0.8439 | 0.7686 | 0.8276 | 2 |
| ————— | ———- | ———- | ———- | ———- | —- |
| s2-s1ep2 | 0.8655 | 0.8441 | 0.7723 | 0.8273 | 3 |
| s2-s1ep4 | 0.8689 | 0.8449 | 0.7668 | 0.8269 | 4 |
| s2-s1ep5 | 0.8706 | 0.8435 | 0.7654 | 0.8265 | 5 |
| s2-s1ep1 | 0.8507 | 0.8382 | 0.7651 | 0.8180 | 6 |
References
- Avsec, Ž. et al. Advancing regulatory variant effect prediction with alphagenome., 649, Nature (2026).
- Avsec, Ž. et al. Effective gene expression prediction from sequence by integrating long-range interactions., 18, Nat. methods (2021).
- Agarwal, V. et al. Massively parallel characterization of transcriptional regulatory elements., 639, Nature (2025).
- de Almeida, B. P., Reiter, F., Pagani, M. & Stark, A. Deepstarr predicts enhancer activity from dna sequence and enables thede novo design of synthetic enhancers., 54, Nat. genetics (2022).
- of Genome Interpretation Consortium, T. C. A. Cagi, the critical assessment of genome interpretation, establishes progress and prospects for computational genetic variant interpretation methods., 25, Genome biology (2024).
- Rafi, A. M. et al. A community effort to optimize sequence-based deep learning models of gene regulation., 43, Nat. biotechnology (2025).
- Lal, A., Garfield, D., Biancalani, T. & Eraslan, G. Designing realistic regulatory dna with autoregressive language models., 34, Genome Res. (2024).
- Alan Murphy, Masayuki Nagai, Alejandro Buendia, Anshul Kundaje, Peter Koo. “Fine-tuning AlphaGenome in native JAX/Haiku.” Genomics × AI Blog, 25 February 2026. https://genomicsxai.github.io/blogs/2026-003/.
Comments
Add your reaction or comment below.