Motivation
AlphaGenome is a foundation sequence-to-function model trained on genome-scale data. Its native JAX/Haiku implementation is powerful but can be cumbersome to modify for custom tasks (we learned this the hard way!). Researchers often want to:
Train a new head on a novel assay
Apply low-rank adapters for efficient backbone updates
Fine-tune the full model progressively
Freeze certain components for stability
Perform attribution analyses to gain insight into learned cis-regulatory logic
To help with this, we developed alphagenome-ft which provides a lightweight wrapper that achieves all these asks without modifying the original AlphaGenome codebase.
Side note: AlphaGenome is a deep learning model that predicts functional genomic signals (e.g., accessibility, transcription, binding) directly from DNA sequence. We call such models sequence-to-function (seq2func) models.
Side note 2: JAX/Haiku are DeepMind’s frameworks which are similar to using PyTorch but optimized for large-scale accelerator workloads.
But wait, why fine-tune AlphaGenome?
Foundation sequence models like AlphaGenome are trained on diverse genome-scale assays, allowing them to learn general regulatory sequence features. However, most research questions involve specific cell types, assays, perturbations, or organisms that differ from the original training distribution.
You should first check if the foundation model alphagenome has an ouput track that’s the same/similar to your cell type of interest, that might be enough! Otherwise, fine-tuning on your cell type/assay of interest is an option.
Fine-tuning adapts the pretrained model to these new contexts while preserving the regulatory knowledge already encoded in the backbone.
Benefits of fine-tuning include:
Improved performance with limited data - Leverages pretrained regulatory features instead of learning from scratch.
Stability and efficiency via frozen parameters - Freezing the backbone while training a new head reduces overfitting, lowers compute cost (this is important AlphaGenome is a BIG model -450m parameters), and prevents catastrophic forgetting.
Parameter-efficient adaptation - Methods such as adapters or partial unfreezing allow targeted updates without retraining the full model.
Faster experimentation cycles - New assays or prediction targets can be incorporated with minimal engineering effort.
Preservation of biological priors - Retains learned sequence motifs and regulatory grammar that remain relevant across assays and cell types.
In practice, many workflows begin by training a task-specific head with the backbone frozen, then progressively unfreezing components if additional capacity is needed.
Fine-tuning with shorter sequence windows
By default, AlphaGenome is trained on ~1 million base-pair (1 Mb) input sequences, allowing the model to capture long-range regulatory interactions. However, during fine-tuning you are not required to use the full 1 Mb context.
If your downstream task does not depend strongly on ultra-long-range interactions, you can fine-tune using shorter input windows (e.g., 32 kb). This reduces memory usage, increases batch size flexibility, and can substantially speed up training.
This is particularly useful when:
The signal of interest is predominantly local (functional outputs like chromatin accessibility are)
You are adapting to assays with shorter effective regulatory range
You want faster experimentation cycles
Importantly, this is different from encoder-only fine-tuning used for very short sequences (e.g., ~200–300 bp MPRA constructs). In that setting, only the convolutional encoder is used, bypassing the transformer and decoder entirely. This is covered in another post.
Here, we are still using the full model stack (encoder → transformer → decoder), but operating on a reduced genomic window.
In practice, reducing input length is a pragmatic trade-off between computational efficiency and long-range regulatory context (i.e. performance).
Package key features
Custom prediction heads – easily register predefined, template, or fully custom heads
Flexible parameter freezing – freeze backbone, individual modules, or heads
Seamless integration – works with pretrained AlphaGenome weights
Parameter inspection – explore and count model parameters
Attribution analysis – gradient-based or in silico mutagenesis (ISM) methods
Native JAX/Haiku – fully compatible with original AlphaGenome pipelines
AlphaGenome fine-tuning workflows schematic
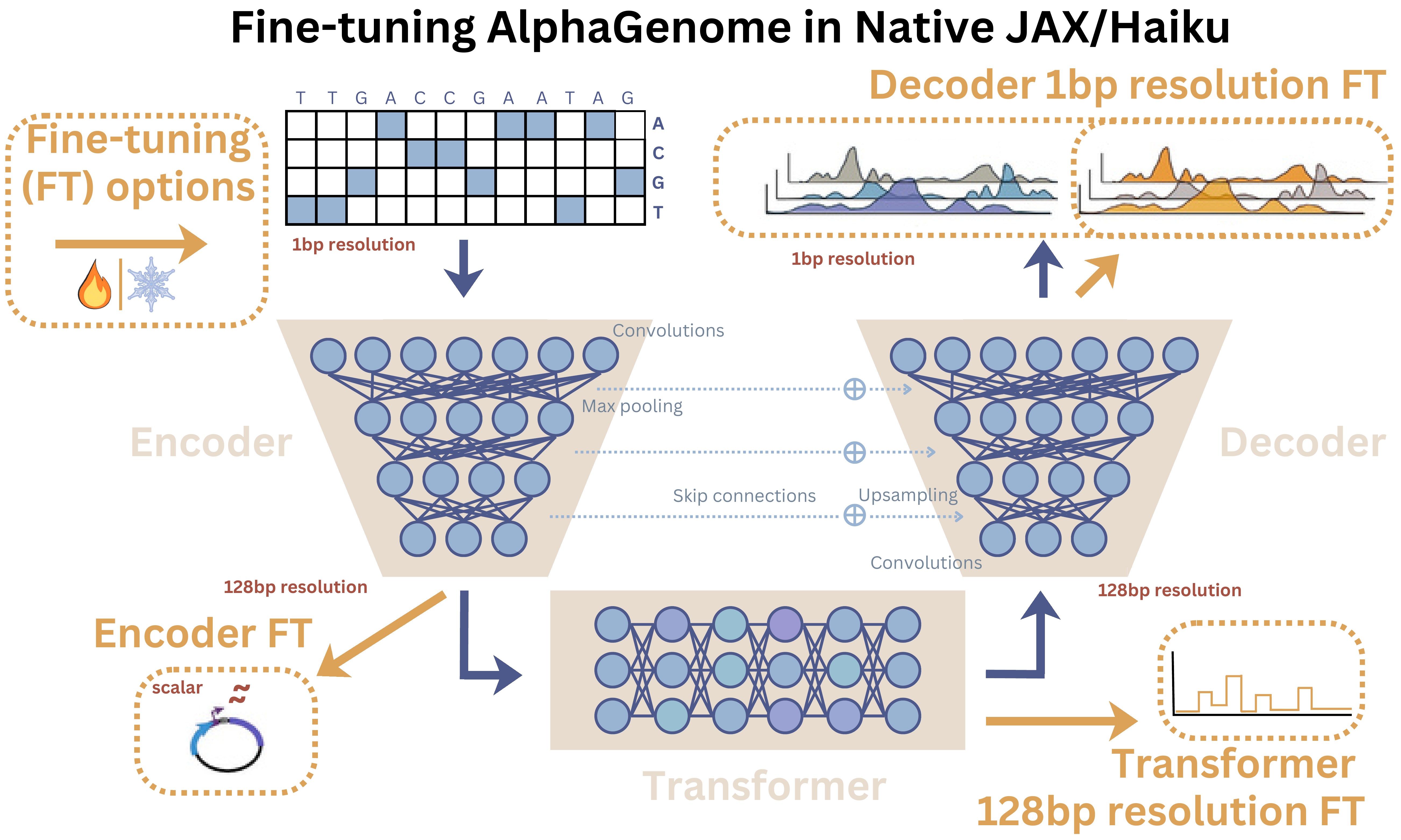
Usage
If these features don’t win you over, let’s walk through how easy it is to use:
Installation
alphagenome-ft wraps AlphaGenome and AlphaGenome Research and is available through pip . Installation requires three steps:
# Step 1: Install AlphaGenome and Research
pip install git+https://github.com/google-deepmind/alphagenome.git
pip install git+https://github.com/google-deepmind/alphagenome_research.git
# Step 2: Install alphagenome-ft
pip install alphagenome-ftPython ≥ 3.11 is required. All other dependencies (JAX, Haiku, optax, etc.) are handled automatically.
Quick Start: Adding new heads
There are two main ways to add heads to AlphaGenome with the package (see the figure above for architecture references):
- Predefined heads
Use existing AlphaGenome head types, e.g., rna_seq, atac, chip_tf:
from alphagenome_ft import (
get_predefined_head_config,
register_predefined_head,
create_model_with_heads,
)
rna_config = get_predefined_head_config("rna_seq", num_tracks=4)
register_predefined_head("K562_rna_seq", rna_config)
model = create_model_with_heads("all_folds", heads=["K562_rna_seq"])
model.freeze_except_head("K562_rna_seq")
#Now ready to train!- Custom heads and reference templates
Our template heads give guidance on accessing different embeddings, which correspond to different biological resolutions - base-pair (bp) precision, regional regulatory context, and short-sequence feature extraction:
StandardHead – 1bp embeddings (decoder output)
TransformerHead – 128bp embeddings (transformer output)
EncoderOnlyHead – CNN encoder output, <1 kb sequences (encoder output)
Note: Template heads are there as a guide for to how to set up your own custom head rather than a definitive ‘best’/‘standard’ option. You should update these with your own layer and loss function choices to fit your data needs.
from alphagenome_ft import templates, CustomHeadConfig, CustomHeadType, register_custom_head
register_custom_head(
'my_head',
templates.StandardHead,
CustomHeadConfig(type=CustomHeadType.GENOME_TRACKS,
output_type='rna_seq',
num_tracks=1)
)Workflows
A full selection of four workflows are given in our github repository, covering:
- Heads-only fine-tuning (frozen backbone)
- LoRA-style adapters (parameter-efficient fine-tuning)
- Full-model fine-tuning
- Encoder-only (MPRA / short sequences)
See the dedicated MPRA post for full post dedicated to Encoder-only fine-tuning (it’s great, even though I may be slightly biased as the one who wrote it …).
Side note: Low-rank adapters (LoRA) enable parameter-efficient fine-tuning by learning small update matrices instead of modifying the full backbone.
If unsure where to start, we recommend training a task-specific head with the backbone frozen, then progressively unfreezing components if additional capacity is needed.
Some extra functionality you might be interested in:
Parameter management and checkpoints
alphagenome-ft allows:
- Modular freezing: encoder, transformer, decoder
- Freezing all heads except one:
model.freeze_except_head('my_head') - Saving checkpoints (heads-only or full model)
- Loading with custom head registration
Attribution analysis
After training, alphagenome-ft also supports:
- DeepSHAP-like attributions - using dinucleotide shuffled reference sequences
- Gradient × Input
- Gradient
- In silico mutagenesis (ISM)
Side note: Attribution methods highlight which nucleotides drive predictions in these models, helping reveal regulatory motifs and sequence grammar learned by the model.
You can also visualise contributions with plot_attribution_map or plot_sequence_logo functions! See below for an example attribution map when we fine-tuned AlphaGenome’s encoder on fly STARR-seq data - See our MPRA post for more details.
 - See our [MPRA post](https://genomicsxai.github.io/blogs/2026-002/) for more details on this fine-tuning. We can see the model highlights an AP-1 motif around position ~230, consistent with known enhancer regulatory logic. attribution map](/blogs/2026-003/sequence_logo_gradient_x_input.png)
You can see we recover the AP-1 motif (TGAsTCA) comes up at roughly position 230 which is a known regulator for developmental genes in flies.
Implications
To take a step back, what do we get with alphagenome-ft? AlphaGenome becomes flexible to:
- Rapid adaptation to new tasks
- Modular freezing/unfreezing for stability
- Supports genome-scale or perturbation assays
- Enables downstream interpretability
AlphaGenome can now be adapted as easily as modern vision and language foundation models — opening the door to rapid regulatory genomics experimentation. So if you think AlphaGenome could be useful if applied to your research, take a look at our package!
alphagenome-ft brings foundation-model-style transfer learning workflows to regulatory genomics.
Compute requirements
- Fine-tuning with a frozen backbone typically fits on a single, high VRAM GPU (e.g., H100).
- Full-model fine-tuning or large batch training will essentially require multi-GPU or TPU setups.
Code and tutorials
- Source code & utilities
- Colab notebooks: Encoder Fine-tuning (MPRA) | Heads-only Fine-tuning
TL;DR
AlphaGenome is a powerful sequence-to-function foundation model, but adapting it natively in JAX/Haiku can be cumbersome.
alphagenome-ft provides a lightweight wrapper for:
adding custom or predefined prediction heads
freezing and unfreezing specific modules
parameter-efficient fine-tuning (e.g., adapters)
running attribution analyses
Most workflows can start by training a task-specific head with the backbone frozen, then progressively unfreezing if needed.
This enables rapid adaptation to new assays while preserving pretrained regulatory knowledge.
If you want to fine-tune AlphaGenome without modifying its core codebase, alphagenome-ft is designed to make that process modular, efficient, and reproducible!
References
- Avsec, Ž. et al. Advancing regulatory variant effect prediction with AlphaGenome., 649, Nature (2026).
- Alan Murphy, Peter Koo. “Adapting AlphaGenome to MPRA data.” Genomics × AI Blog, 20 February 2026. https://genomicsxai.github.io/blogs/2026-002/
- Hu, E. J. et al. Lora: Low-rank adaptation of large language models (2021), https://arxiv.org/abs/2106.09685. 2106.09685.
- de Almeida, B. P., Reiter, F., Pagani, M. & Stark, A. Deepstarr predicts enhancer activity from dna sequence and enables the de novo design of synthetic enhancers., 54, Nat. genetics (2022).
Comments
Add your reaction or comment below.